PyMOL
What is PyMOL?
PyMOL is an open source molecular graphics system with an embedded Python interpreter designed for both real-time visualization of 3D molecular structures and generation of high-quality renderings and animations. Having been created by Warren Lyford DeLano, PyMOL was initially commercialized by DeLano Scientific LLC, but is currently marketed by Schrödinger LLC. For more details, please consult either the Home page of incentive PyMOL by SCHRÖDINGER or the Home page of the open source project on GitHub.
Usage of PyMOL at LRZ
The environment modules package controls access to the software. Use "module avail py" to find all available versions of PyMOL installed at LRZ. The output could look as follows:
~> module av py ---------------------------------------- /lrz/sys/spack/release/21.1.1/modules/x86_64/linux-sles15-x86_64 --------------- py-pymol/2.3.0 python/3.7.10-base python/3.7.10-extended python/3.8.8-base python/3.8.8-extended ---------------------------------------- /lrz/sys/share/modules/files_sles15/tools --------------------------------------
From here you can learn, that the current version of PyMOL in SPACK software distribution system is obviously named "py-pymol/2.3.0". To use the default version of PyMOL, please type:
> module load py-pymol/2.3.0
Visualization using the PyMOL GUI
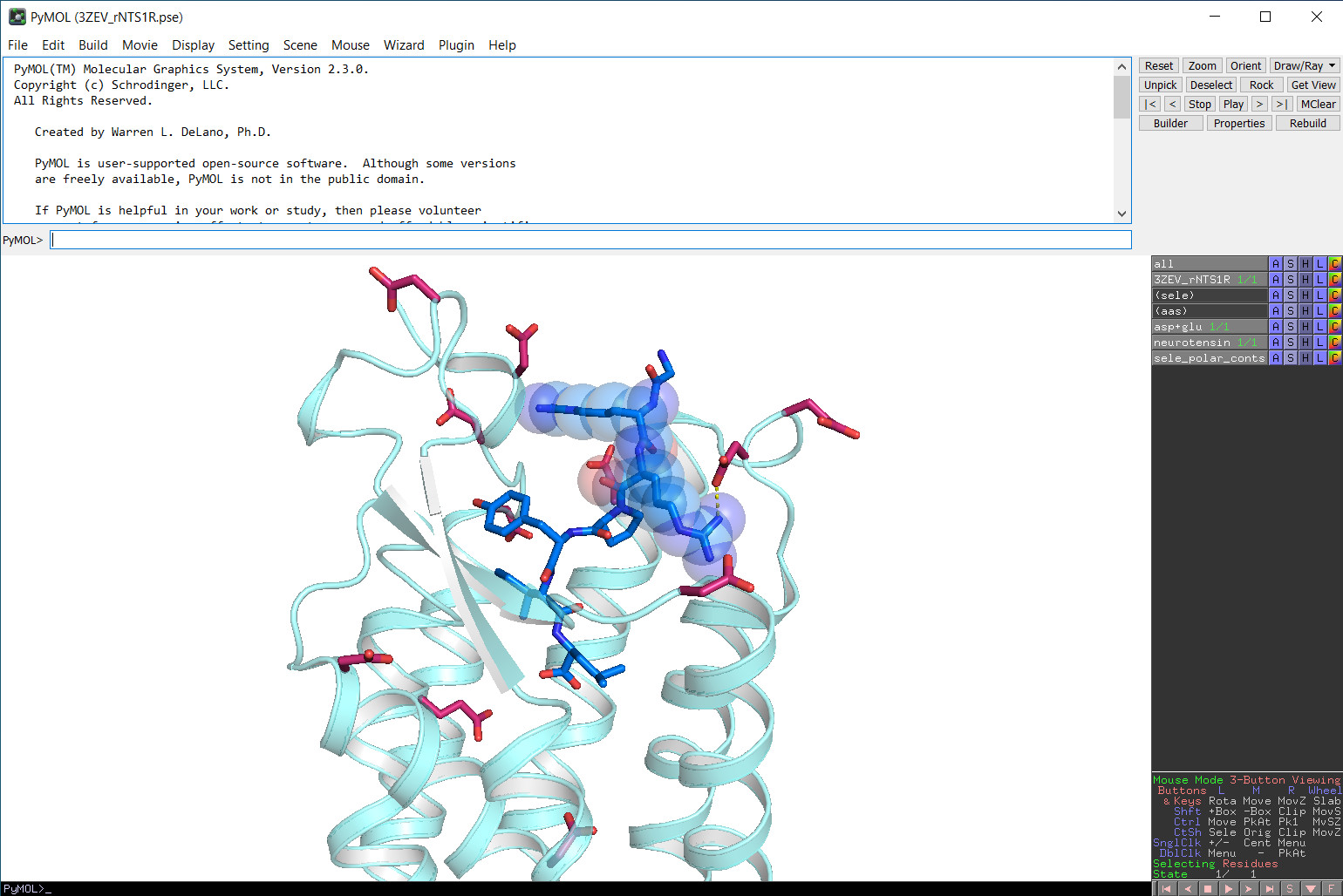
You can start the PyMOL GUI either on the login nodes of Linux-Cluster & SuperMUC-NG or by using our remote visualization service (via VNC). We however want to note that the graphics performance on the login nodes is comparatively poor. Therefore, we recommend our users to only start the PyMOL GUI on the login nodes for less demanding visualisations of e.g. small molecules, but not for CPU-intensive renderings using "ray". If you however want to either visualize larger data sets or perform CPU-intensive renderings using "ray", we recommend using our remote visualization service (via VNC) instead. You will then entirely have access to the reserved "localhost" node including GPU resources and can run CPU-intensive tasks within the PyMOL GUI.
To start PyMOL graphical user interface, simply type:
> pymol
Running PyMOL on the command line (for productive-level, publication-ready renderings)
Given the fact that rendering of 3D molecular structures can be very CPU-intensive tasks, we recommend our users to run PyMOL via the command line (salloc/srun or sbatch) on a whole node of either Linux Cluster or SuperMuc-NG to prepare high-quality, publication-ready molecular images.
For this purpose, you need a previously generated (using the PyMOL GUI either as shown above or on your local computer) ".pse" file containing both 3D coordinates of the molecular structures and information on how the structures should be visualized (e.g. cartoon or ball-and-stick representation). Furthermore, you need a ".pml" file (for example file see below) containing at least the "load" command to import the ".pse" file, the "ray" command with the number of pixels in width and hight to render/raytrace the molecular structures and the "png" command to save the rendered scene in a ".png" file. Furthermore, the ".pml" file can contain additional rendering commands (e.g., set_view ..., space rgb/cmyk, hide ..., set ...) that at least in part could have been applied at the time the ".pse" file has been generated.
load PSE_FILE_NAME.pse ray 2000, 1500 png PNG_FILE_NAME.png
Via an interactive SLURM shell (recommended for a few jobs or testing purposes)
To e.g. request one node on mpp2 of the Linux Cluster for half an hour, please issue:
> salloc --nodes=1 --time=00:30:00 --partition=cm2_inter
For requesting one node on the test partition of SuperMuc-NG for half an hour, please issue:
> salloc --nodes=1 --time=00:30:00 --partition=test --account <project id>
When a new batch session is returned to you, simply type
> module load py-pymol/2.3.0 > srun pymol -c PML_FILE_NAME.pml
to render/raytrace the scence and save it in a ".png" file.
By submitting a SLURM batch script (recommended for production runs)
For production runs, please create a SLURM batch script and submit via "sbatch".
Linux Cluster | SuperMUC-NG |
|---|---|
#!/bin/bash #SBATCH -o /dss/dsshome1/<group>/<user>/mydir/%x.%j.out | #!/bin/bash |
Documentation
For documentation, please consult either the incentive PyMOL Documentation Home page, the PyMOL Command Reference or the Quick Reference Guide for Intermediate PyMOL Users.
Support
If you have any questions or problems with PyMOL installed at LRZ platforms, please don't hesitate to contact LRZ HPC support staff.